|
This has led to the development of methods for quantifying cell fusion independent of microscopic inspection. Furthermore, as fusion index determinations are generally carried out manually, they are laborious, error-prone and often inaccurate. In addition, cells growing on top of each other can be mistaken for syncytia. However, without continuous monitoring, it is impossible to decide by microscopy alone whether multinucleation is caused by cell fusion or the result of karyokinesis without cytokinesis. Ĭell fusion can be easily observed using microscopic techniques and in many studies the extent of cell fusion is expressed as fusion index, which either stands for the percentage of cells with two or more nuclei or the percentage of nuclei present in syncytia. Inappropriate cell fusion has been implicated in tumor development and progression. 
In general, cell fusion is a tightly regulated and highly selective process involving specific cell types. Plasma membrane fusion is a crucial event during, for example, fertilization, syncytiotrophoblast production, skeletal muscle formation, bone remodeling, eye lens development and certain forms of tissue repair. 
The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.Ĭompeting interests: The authors have declared that no competing interests exist.ĭuring cell-to-cell fusion, plasma membranes of individual cells merge to form a multinucleated structure called a syncytium. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.įunding: This work was supported by research grants from the Dutch Duchenne Parent Project (DPP) and the Iranian Ministry of Science, Research and Technology. Received: JAccepted: JPublished: July 16, 2014Ĭopyright: © 2014 Neshati et al. PLoS ONE 9(7):Įditor: Atsushi Asakura, University of Minnesota Medical School, United States of America In summary, a rapid and simple chemiluminescence assay for quantifying cell-to-cell fusion progression based on GpLuc has been developed.Ĭitation: Neshati Z, Liu J, Zhou G, Schalij MJ, de Vries AAF (2014) Development of a Lentivirus Vector-Based Assay for Non-Destructive Monitoring of Cell Fusion Activity. However, at high FLPe concentrations the presence of the NLS negatively affected reporter gene expression. The ability of FLPe NLS+ to spread through myofibers and to induce reporter gene expression is thus not limited by its NLS. 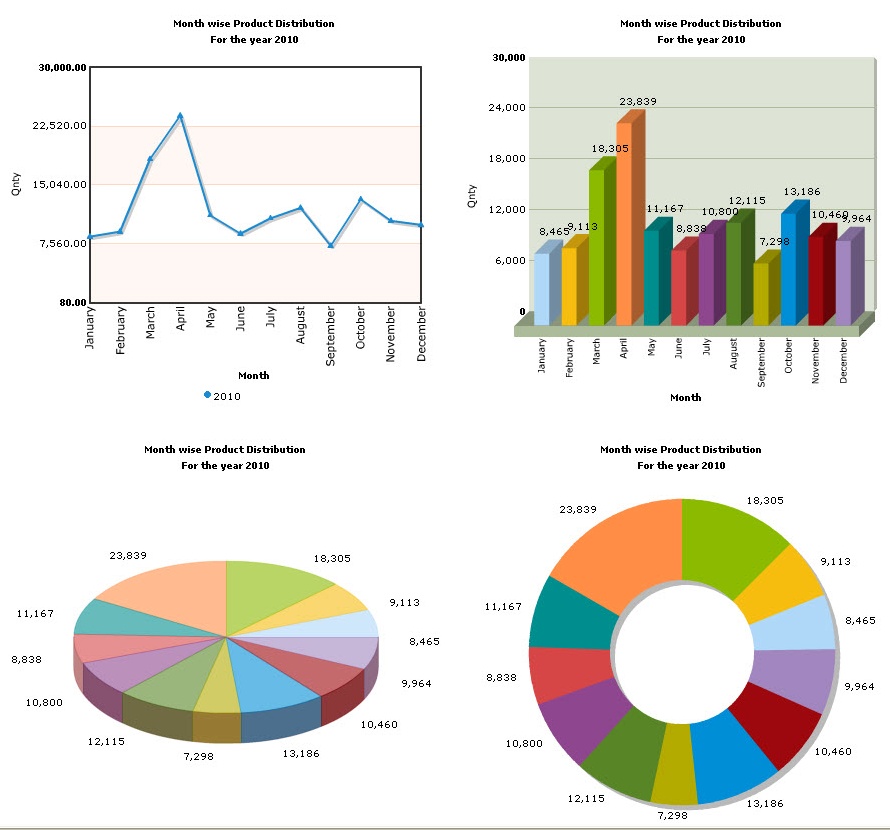
FLPe NLS+ and FLPe NLS− both activated the latent GpLuc gene but when the percentage of FLPe-expressing myoblasts was limiting, FLPe NLS+ generally yielded slightly higher signals than FLPe NLS− while at low acceptor-to-donor cell ratios FLPe NLS− was usually superior. At different times after induction of cell-to-cell fusion the GpLuc activity in the culture medium was determined. 
To test this hypothesis, myoblasts were transduced with LVs encoding either FLPe NLS+ or an NLS-less version of FLPe (FLPe NLS−) and subsequently co-cultured in different ratios with myoblasts containing the FLPe-activatable GpLuc expression cassette. In myotubes the spread of FLPe NLS+ may be limited due to its nuclear localization signal (NLS) causing low signal outputs. Therefore, in this study the PpLuc-coding sequence was replaced by that of Gaussia princeps luciferase (GpLuc), a secretory protein allowing repeated analysis of the same cell culture. PpLuc activity is typically measured in cell lysates, precluding consecutive analysis of one cell culture. Fusion of both cell populations will lead to the FLPe-dependent generation of a functional PpLuc gene. One way to accomplish this goal is by using a bipartite lentivirus vector (LV)-based cell fusion assay system in which the cellular fusion partners are transduced with a flippase-activatable Photinus pyralis luciferase ( PpLuc) expression unit (acceptor cells) or with a recombinant gene encoding FLPe NLS+, a nuclear-targeted and molecularly evolved version of flippase (donor cells). Cell-to-cell fusion can be quantified by endowing acceptor and donor cells with latent reporter genes/proteins and activators of these genes/proteins, respectively.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
